To colonize the nasal cavity, bacteria must attach to the epithelial surface, accommodate or evade the immune system, and extract minute nutrients and minerals from their host. Together, these features make the nasal cavity a hostile and competitive environment. As one means to persist in this system, bacteria engage in competitive interspecies interactions.
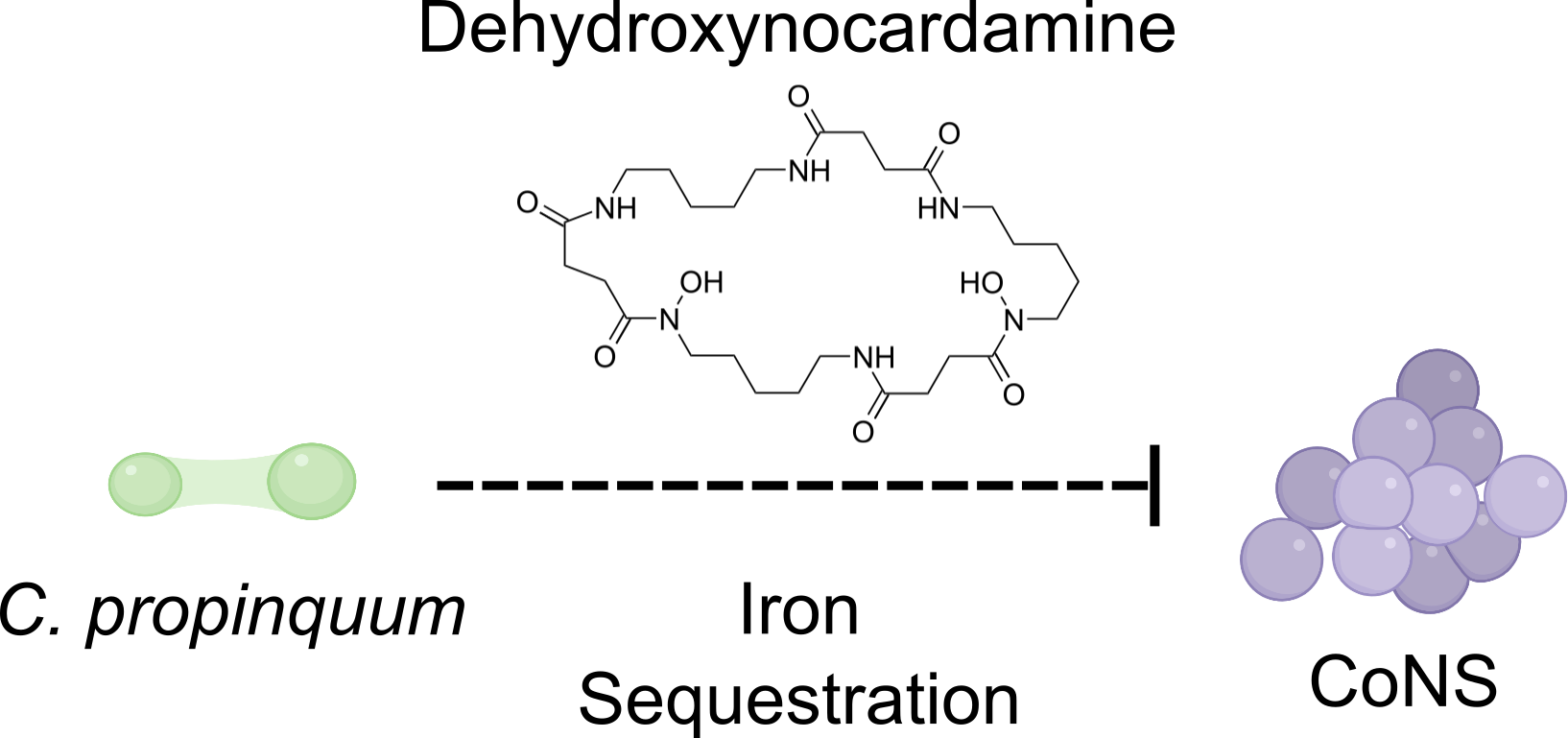
Iron is a limiting resource in the nose. In previous work, we determined that Corynebacterium propinquum strains deplete the local environment of iron using the siderophore dehydroxynocardamine, which inhibits the growth of competing coagulase-negative staphylococci (CoNS). By mining metatranscriptomic reads from the NIH Human Microbiome Project, we were able to determine that the dehydroxynocardamine biosynthetic gene cluster is expressed in vivo. This work provided the first evidence for resource competition in the nasal cavity.
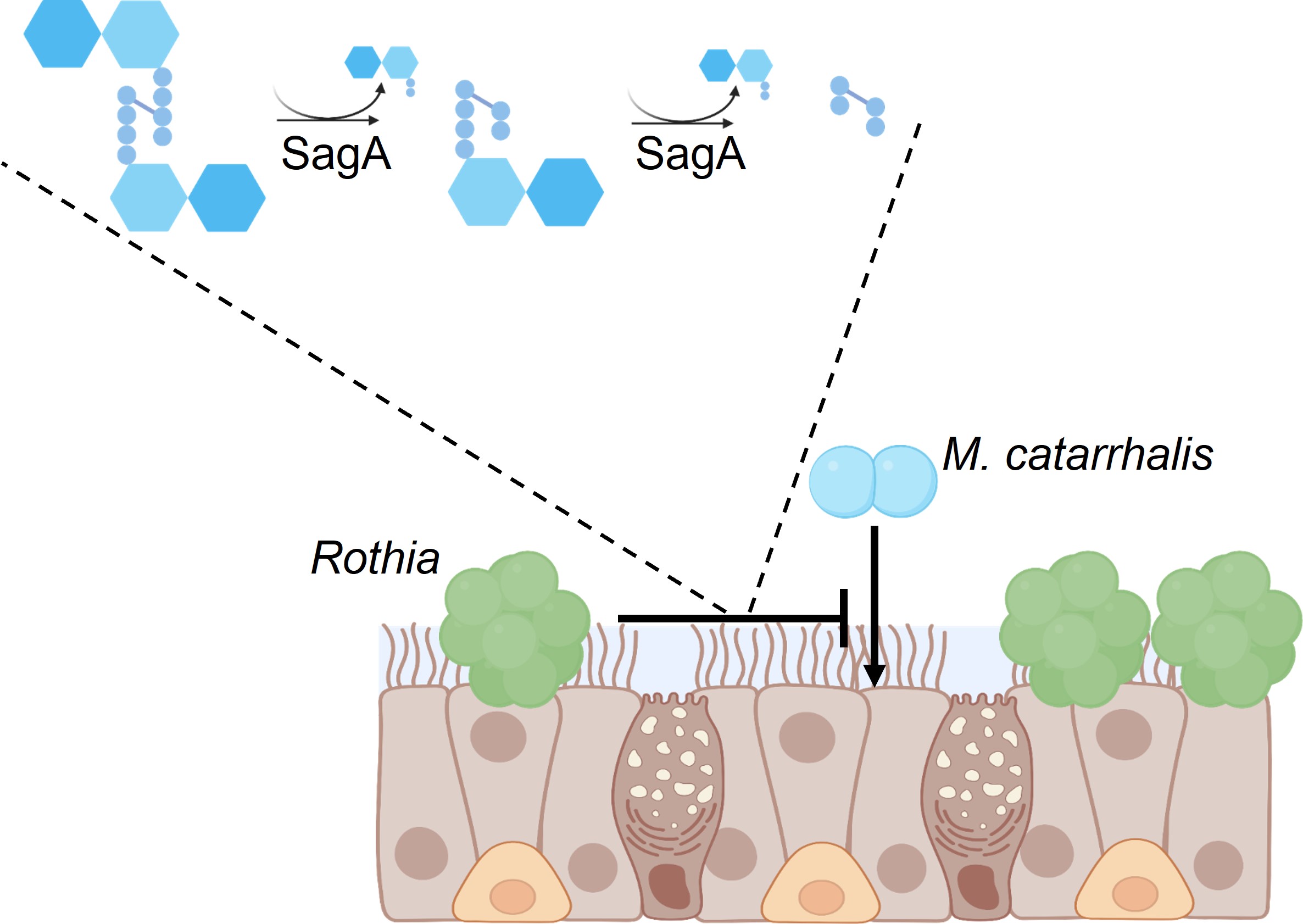
The human nose is colonized by opportunistic pathogens, including Moraxella catarrhalis. This pathobiont is found almost exclusively within the human respiratory tract and is commonly isolated as a causative agent of acute otitis media. In addition, this pathobiont has become recently associated with the development of respiratory diseases, including asthma and allergy. In previous work, we identified that Rothia was more abundant in the noses of healthy children without M. catarrhalis than the noses of sick children. By characterizing the secreted proteomes of Rothia, we discovered that these bacteria produce an endopeptidase that degrades M. catarrhalis peptidoglycan called secreted antigen A (SagA). In collaboration with James Gern (University of Wisconsin, Departments of Pediatrics and Medicine), we showed that Rothia reduced M. catarrhalis levels in an air-liquid interface culture model of respiratory epithelium.
Future studies in the laboratory will focus on identifying additional metabolites and proteins that are involved in commensal-commensal and commensal-pathogen interactions in the nose.